QIAGEN CLC Genomics
RNA-seq and single-cell RNA-seq data analysis using QIAGEN CLC Genomics Workbench_Sep 22
2,483 views
For RNA-seq and scRNA-seq data, users will learn how to:
• Import fastq files, cell matrix files and metadata and how to download references
• Map reads to a reference genome and generate gene and transcript counts and QC reports displaying % mapped reads, knee plots, etc.
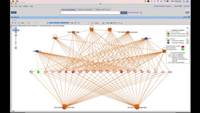
• Generate visualizations of results, such as heatmaps, differential expression tables, PCA/PCOA plots, t-SNE plots, UMAP plots, dot plots, Venn diagrams and others
• Annotate single-cell clusters overlain with gene expression
• Easily customize RNA-seq and scRNA-seq workflows
• Export publication-quality graphics, tables and reports
• Send differential expression table to QIAGEN Ingenuity Pathway Analysis directly from QIAGEN CLC Genomics Workbench
Speaker: Araceli Cuellar, Ph.D., Field Application Scientist, QIAGEN Digital Insights
Related videos
QIAGEN CLC Genomics
QIAGEN CLC Main Workbench training
Day 1: QIAGEN CLC Main Workbench Speaker: Shawn Prince, Sr. Field...
QIAGEN CLC Genomics
Biologically interpret your RNA-seq data with knowledge-based IPA
This webinar will provide an overview of how to analyze data obtained from...
QIAGEN CLC Genomics
QIAGEN CLC Genomics Workbench 20 – scalable desktop software for NGS data analysis
In this webinar, Leif Schauser, Ph.D., Director Product Management Genome...
QIAGEN CLC Genomics
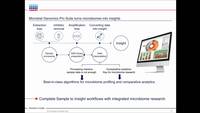
Data analysis on day one - Taxonomic and functional microbiome profiling with the Microbial...
In this webinar, we will introduce the scientist-friendly Microbial Genomics...