QIAGEN CLC Genomics
Isolate typing, strain identification and antimicrobial resistance analyses using QIAGEN CLC Genomics Workbench - Aug 25 2021
850 views
Topics covered in this 90-minute webinar include:
• An overview of the different tools within QIAGEN CLC Microbial Genomics Module and the research areas supported
• A focused review of isolate typing and characterization, including:
o Importing data
o Utilization of metadata
o Downloading and managing references
• Database of isolates/ resistances/ multi-locus sequence typing (MLST) analysis
o A walk-through of the ‘Type a Known Species’ workflow
- Review details for each isolate
o Creating SNP profiles to a specific reference
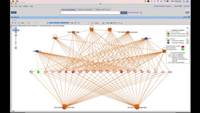
o Generate a SNP tree for isolate comparison
o Export tabular and high-quality graphical outputs in a wide range of file formats
Speaker: Shawn Prince, Ph.D., Field Application Scientist, QIAGEN Digital insights
Related videos
QIAGEN CLC Genomics
QIAGEN CLC Main Workbench training
Day 1: QIAGEN CLC Main Workbench Speaker: Shawn Prince, Sr. Field...
QIAGEN CLC Genomics
Biologically interpret your RNA-seq data with knowledge-based IPA
This webinar will provide an overview of how to analyze data obtained from...
QIAGEN CLC Genomics
QIAGEN CLC Genomics Workbench 20 – scalable desktop software for NGS data analysis
In this webinar, Leif Schauser, Ph.D., Director Product Management Genome...
QIAGEN CLC Genomics
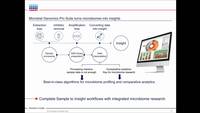
Data analysis on day one - Taxonomic and functional microbiome profiling with the Microbial...
In this webinar, we will introduce the scientist-friendly Microbial Genomics...